We have found three bacteria that are capable of decomposing PETs. Pseudomonas has 6.3 million base pairs which makes it one of the longest sequenced bacterium. We had difficulty finding the genetic sequences of our bacterium and enzymes.
0 Comments
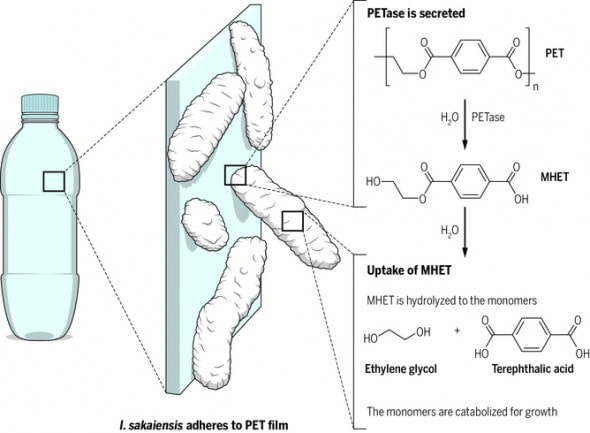
During further research, we discovered a bacteria that is called Ideonella sakaiensis this bacteria dissolves plastics using different enzymes. This is good because this bacteria is easy to find, and can be reproduced quickly and easily which means that it could be a successful solution to the plastic pollution crisis. Researchers have looked into this bacteria and performed an experiment in which they put the bacteria inside of a jar with PET along with a few nutrients and after a few weeks the PET that was inside of the jar was completely gone.The bacterium uses PETase an enzyme that works with water to break down PET plastic, then uses MHETase to break down PET even more, MHETase is another enzyme that also reacts with water to break down the plastics into terephthalic acid and ethylene glycol two substances that do not pose a threat to the environment.While doing research, we discovered worms that are able to digest plastics. This is very special because it can help the research behind the biodegradation of plastic. which is very important because people use about 300 million tons of plastic every year and a large amount of it ends up in the ocean causing many environmental issues. We are looking to find what enzyme inside of these worms are able to break down these plastics so that we may use it to lower the amount of plastic in the environment. This enzyme is great because it breaks down the plastic at a molecular level and leaves behind no waste.We have been studying the many different kinds of plastics found in the world's waterways. We hope to find a solution to the pollution problem using synthetic biology. The goal is to find/create an organism that can break down plastics into materials that are not harmful to the environment
|
Authors
Rafael Faria Archives
|

 RSS Feed
RSS Feed
